
Biosignal processing comes to the Web
Building applications with biosignals on the web has historically meant building everything from scratch. From BLE (Bluetooth Low Energy) device transports, signal processing pipelines, and protocol decoders, one thing is clear: there is no universally standardized workaround.
In fact, in brain sciences, most tooling still assumes MATLAB and a six-figure hardware budget.
For builders, this means that if you want neural signal data or contactless biosignal acquisition, you've had to piece it together yourself. This arduous process would typically include porting research code, fighting web Bluetooth APIs, and settling for slow JavaScript implementations of algorithms that need real performance.
Clearly, this is an area that has not yet caught up to 21st century technology standards.
With Elata SDK, we're enabling a new class of applications built on top of anonymous neural signaling that are fully abstracted. You don't need to even have expensive hardware to use apps. In fact, you don't need even hardware at all; apps can be used contactlessly through your phone or computer's native camera.
Why the web
The browser is the most universal application runtime on earth. It's cross-platform by default, requires no installation, and reaches anyone via. a URL.
However, biosensing has never lived on the web. Traditionally, the computation has been too heavy for JavaScript. The device protocols are too low-level for most web developers, so the signal processing expertise is concentrated in academic labs, not npm packages.
Elata SDK solves this by doing the hard work in Rust, compiling to WASM, and exposing it through TypeScript packages that feel like any other npm dependency.
The performance is intended to be near-native. The developer experience is fully abstracted and intended to accommodate developers of all skillsets: it's just npm install, and build.
Because everything runs client-side in WASM, raw sensor data never has to leave the device. The browser isn't just the delivery mechanism, but rather, the processing environment.
The architecture
As this is the initial release of the Elata SDK, our EEG packages work strictly with Muse headsets, most notably the Muse Athena S and the Muse 2.
Currently the Elata SDK is comprised of three packages, each targeting a different sensing modality:
@elata-biosciences/rppg-web processes webcam video to extract biosignals. Heart rate is detected through remote PPG signals collected, plus facial affect and sentiment analysis. No hardware is required at any point in time for this modality.
@elata-biosciences/eeg-web handles EEG signal processing. Band power extraction across all standard frequency bands (delta, theta, alpha, beta, gamma), FFT, filtering, and two built-in analysis models: an Alpha Bump Detector that tracks relaxed vs. alert state transitions, and a Calmness Model that outputs a continuous 0-100% score from the alpha/beta power ratio.
@elata-biosciences/eeg-web-ble is the Web Bluetooth transport layer for HAL-compatible EEG headbands. It handles device discovery, BLE pairing, and protocol decoding, then emits clean, normalized frames that pipe directly into eeg-web for processing.
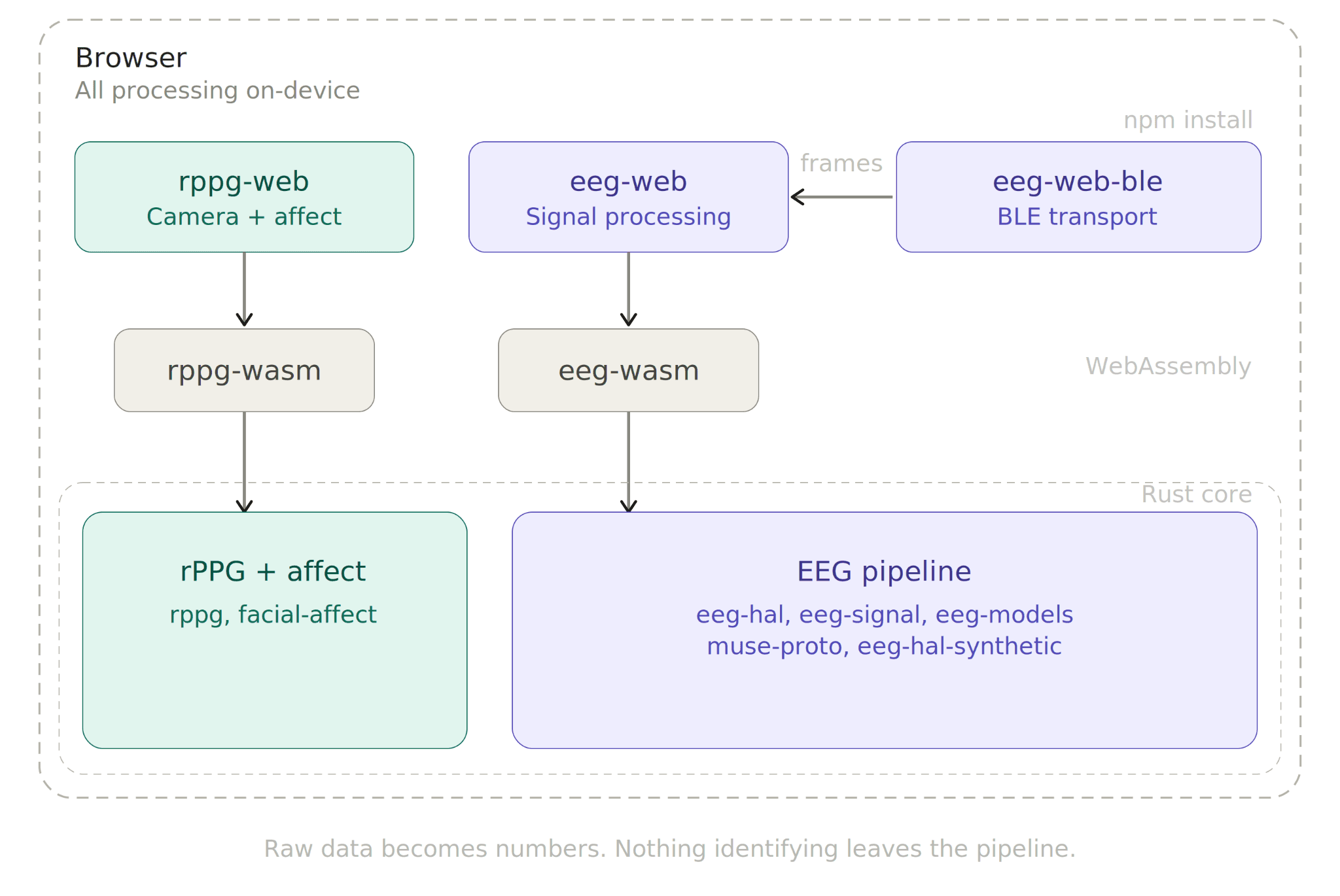
Together, the packages cover contactless biosensing (camera), neural signal physiology (EEG), and behavioral analysis, all from the browser, all from npm install.
How it works
Elata SDK's core is written in Rust and organized as a workspace of fourteen crates spanning the signal pipelines mentioned in the section prior. Rust compiles to WebAssmebly (WASM) for the browser and to native libraries for iOS, Android, and desktop via UniFFI. The TypeScript packages wrap the WASM bindings with ergonomic APIs.
A few things worth noting:
Hardware Abstraction Layer (HAL): The EEG pipeline is built on HAL traits (EegDevice, SampleBuffer, ChannelConfig).
A particularly amazing and unique thing about Elata SDK is that it's open-source, making it such that contributors can plug any at-home neural sensing device into the SDK by implementing these traits.
For the initial release, the HAL works with Muse's line of headbands and a synthetic device for testing. The interface is open for any hardware to follow.

Synthetic device for development: eeg-hal-synthetic provides a configurable synthetic EEG source with signal profiles (relaxed, alert, etc.). You can build and test your entire application without owning a device.
You can build now, ship to people who already have hardware, and then swap to real hardware when you're ready to use it yourself.
WASM-optimized builds: The release profile uses opt-level = "s" with LTO, explicitly tuned for small binary size. This is a browser-first SDK. Bundle size matters and we treat it that way.
One codebase, every platform: The Rust core compiles to WASM for browsers, native Swift for iOS, native Kotlin for Android, and runs directly on desktop. You can write signal processing once, and ship everywhere.
Elata SDK never sees a person. It sees signals. All processing happens locally on-devices. Raw sensor data, whether camera frames or EEG samples, is literally just reduced to physiological metrics in the browser.
What comes out the other end is a number, not a biometric. A band power reading or a heart rate value carries no identity. Identity is lost in the pipeline because the pipeline was never looking for it. Nothing leaves the browser unless the developer explicitly decides it should.
Getting started
Full API documentation and usage examples are in the repo README and each package's source directory.
What you can build
The entire point of us porting this capability to the web and consumer-facing areas is for biosignal and brain-interfacing applications to be completely ubiquitous.
We want to lower the barriers to entry and enable web and application-facing developers to build and ship biosignal apps that they previously couldn't. This could include a ton of things, and since processing occurs on user devices, can include incredibly unique use-cases such as brain-compatible AI applications, games, biometric analysis apps, and so on. Here are a couple more precise examples:
A browser-based meditation app with live EEG biofeedback. Connect a headband via Web Bluetooth, stream alpha/beta powers, and drive a visual or audio feedback loop.
A stress monitor that runs on your camera. Track HR, HRV, and facial micro expressions over a work session with no wearables at all through a mobile app or the web (i.e. browser extension or website).
A neuroscience experiment that runs entirely in Chrome. Simply recruit participants via a link. They grant camera or Bluetooth access, complete the protocol, and you collect derived metrics, not raw biometric data. No installation, no IT department, no lab visit, and no sensitive data pipeline to secure.
A game where cognitive state drives gameplay. Pipe alpha state detection into game mechanics. Relaxation charges a power meter. Focus sharpens aim. The input device is your brain.
These are not future plans. They are buildable today, with the packages that are published right now.
Having brought the barrier to entry so low, builders can simply use modern AI coding tools to build apps in incredibly short periods of time.
Build with us
Elata SDK is open source under MIT. Anyone can use it.
In particular (and since this is the initial release), we're looking for:
Builders who will install these packages and ship something. We want to see what happens when biosensing is as easy as importing a library.
Rust contributors who want to add device drivers, signal processing algorithms, or new analysis models to the core.
Researchers who need browser-based experiment tooling and are done waiting for the MATLAB-to-native-app pipeline to modernize.
Device makers who want their hardware to work with an open ecosystem rather than a proprietary SDK that only they maintain.
Here's where to start:
Contributor guide: CONTRIBUTING.md
Documentation: docs.elata.bio
The infrastructure for biosensing on the web didn't exist before. Now it does. Open source, on-device, non-identifying by design. What will you build?





